Introduction
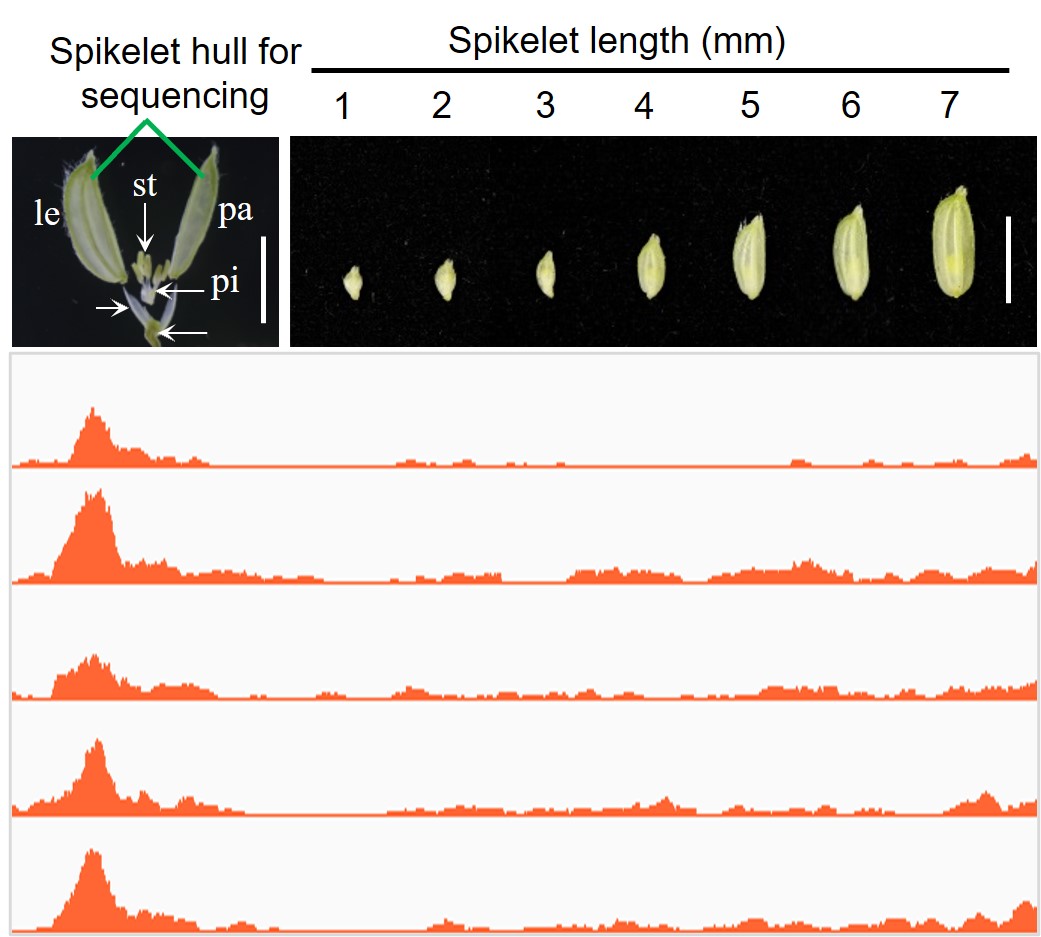
We performed time-course RNA-seq and ATAC-seq to characterize the transcription and chromatin accessibility dynamics during the development of spikelet hull in the indica rice Minghui 63 (MH63). We found a sequential expression pattern of more than half of expressed genes, in which related pathways are organized hierarchically to regulate gain size development. By targeting the open chromatin regions upstream of the start codon of genes, it is possible to fine-tune the grain size.
Here, we provide a visualization of the RNA-seq profiles and ATAC-seq data. You can view the data in Genome Browser function, including the covered reads and ATAC-seq peaks around a gene. The genome is based on the MH63RS3 assembly. For convenient, we provide a mini-tool (ID conversion) for users to convert the gene ID of Nipponbare reference (LOC or OS) to the ID of MH63RS3.